Long in the tooth: Ancient proteins illuminate deep-time species evolution
07 March 2023 | Story Helen Swingler. Read time 6 min.
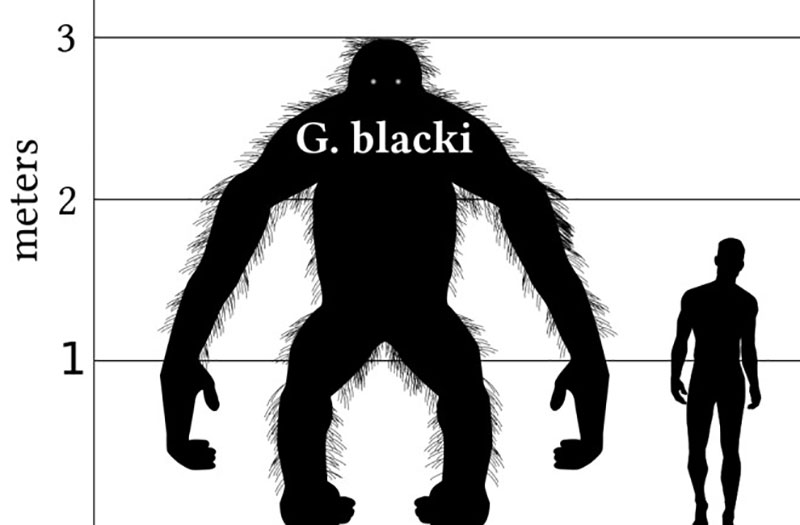
Standing up to 3 m tall and weighing up to 600 kg, illustrations of the massive primate Gigantopithecus blacki call to mind the mythical Himalayan Yeti. Dental remains of the prehistoric ape, which lived in the tropical forests of south China, have been dated to 1.8 million years ago, and scientists now know they were ancient relatives of modern-day orangutans. This is thanks to advances in palaeoproteomics, keys to a vault of significant palaeontological data.
Broadly, palaeoproteomics is the study of ancient proteins. These are ‘mined’ from mineralised fossil remains such as bone, teeth, and eggshell. And the example of Gigantopithecus blacki was one of several presented at a public seminar titled “Reconstructing Deep-Time Evolution Using Palaeoproteomics”.
This public seminar formed part of the African Palaeoproteomics Workshop, held at the University of Cape Town (UCT) from 23 to 25 February, and hosted by the university’s Human Evolution Research Institute (HERI).

The guest speakers were Associate Professor Enrico Cappellini and PhD candidate Palesa Madupe of the Globe Institute at the University of Copenhagen. Madupe is affiliated with HERI, and is co-supervised by HERI co-director, Professor Rebecca Ackermann. A pioneer in the field, it was Associate Professor Cappellini’s second visit to UCT.
Game-changing techniques
Proteomics is not new, having developed after the advent of soft-ionisation mass spectrometry-based protein sequencing in the United States in the late 1980s and early 1990s. But modern palaeoproteomics is new and has been a game-changer for scholars of evolution and taxonomy.
“The field of palaeoproteomics is relatively young and African researchers, particularly young African researchers, can contribute to it immensely.”
“The field of palaeoproteomics is relatively young and African researchers, particularly young African researchers, can contribute to it immensely and should be given the opportunity to do so,” said Madupe.

Palaeoproteomics allows researchers to ‘see’ further back in time than ancient DNA (DNA degrades too quickly), to investigate much older species and to map out deep-time evolution. Sequencing ancient DNA enables scientists to reconstruct speciation, migration and admixture events (when individuals from two or more previously isolated populations interbreed) for these extinct taxa.
However, the irreversible postmortem degradation of ancient DNA has so far limited its recovery – outside permafrost areas where frigid conditions help to preserve fossils – to specimens that are not older than approximately 0.5 million years.
Cappellini and his team have shown that proteins recovered from human and primate fossils up to about 2 million years old can be reliably used to confidently reconstruct the evolutionary relationships between extant species and the extinct ones whose DNA is completely lost.
In addition to using palaeoproteomics to reconstruct the evolutionary history of Gigantopithecus blacki, Cappellini and colleagues have also used this approach to understand the evolutionary placement of a 1.7-million-year-old species of woolly rhinoceros, and the extinct European hominin species Homo antecessor.
Dragon teeth
The story of Gigantopithecus blacki offers a fascinating glimpse of what tiny amounts of fossil protein can reveal. The massive primate is thought to have gone extinct at the beginning of the last Pleistocene ice age when climate change transformed its densely forested habitat into savannah grasslands, a diet unsuited to the large frugivore.
The fossil record for Gigantopithecus blacki was scant; no skeletal material, only partial jawbones, fortunately with many impressively large teeth. These even found their way into the commercial arena. A 2019 article in the Smithsonian Magazine recalls that Western scientists first came across Gigantopithecus blacki in 1935 when an anthropologist found its huge molars in Chinese drug stores. These were reportedly sold as dragons’ teeth.
But it was a molar later found in a cave in China, where four jawbones were recovered, that finally helped Cappellini and colleagues to determine its closest living relative: the orangutan. Protein was extracted from the dental enamel and analysed using mass spectrometry.
What can this tell us about us?
Madupe’s PhD work concerns palaeoproteomic investigations of South African hominins, such as the species Paranthropus robustus. The South African fossil record is vital for our understanding of human evolution in Africa – from early australopiths to the origin of our species. These methods allow us to determine biological sex of individuals, as well as to produce trees of species relationships.

Participants in the NRF AOP-funded African Palaeoproteomics Workshop. Photo Lerato Maduna.
Results from her work have already been presented at a conference at the European Society of Human Evolution and are due for publication soon.
In South Africa possibilities are beginning to mushroom, tapping into a growing network of countries and institutions involved in the field.
That was the goal of the African Palaeoproteomics Workshop, funded through the National Research Foundation’s African Origins Platform with ring-fenced African Engagements funding, said Professor Ackermann.
“Our workshop was an exciting, invigorating space. We are thrilled to take this research forward and to formalise a network.”
The workshop brought together more than two dozen researchers and promising students from South Africa, Kenya, Tanzania, and Ethiopia, in addition to African colleagues in the diaspora, for exciting collaboration exploring future pan-African palaeoproteomics research directions.
This forms part of HERI’s wider goal to create a pan-African training network.
Ackermann said, “Our workshop was an exciting, invigorating space. We are thrilled to take this research forward and to formalise a network that supports young, emerging African talent in human evolution research.”
 This work is licensed under a Creative Commons Attribution-NoDerivatives 4.0 International License.
This work is licensed under a Creative Commons Attribution-NoDerivatives 4.0 International License.
Please view the republishing articles page for more information.
Research & innovation





































