Worldwide brain study is boosted by NIH grant
14 October 2014 | Story by Newsroom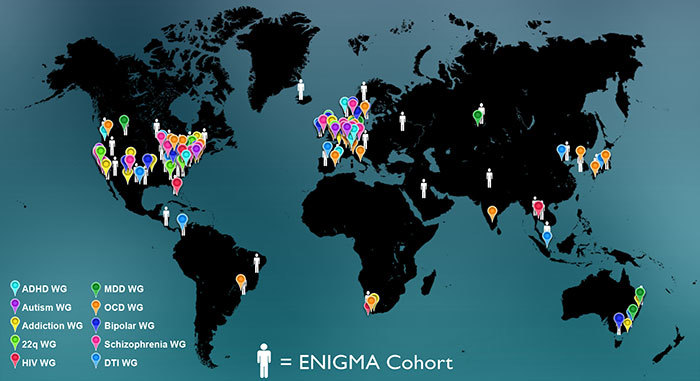
A global initiative to pool data about the human brain has been funded by the US National Institutes of Health (NIH). Project ENIGMA, named after an allied code-breaking initiative in World War II, unites brain researchers in 33 countries to discover factors that help or harm the brain. The research alliance studies medical scans of the brain and DNA collected from 30 000 people.
"Our effort brings a new source of power to biomedical science: our global alliance pools brain scans and DNA in a monumental global effort to understand brain diseases," says Paul Thompson, co-founder of ENIGMA, and a professor of neurology at the University of Southern California in Los Angeles. "The combined data is so vast that no one scientist could ever analyse it on their own. Together, we can screen each other's data to discover genes or medications that influence a range of neuropsychiatric disorders."
 Dan Stein, head of the Department of Psychiatry at the University of Cape Town.
Dan Stein, head of the Department of Psychiatry at the University of Cape Town.
"The ENIGMA project gives us a power we have not had," says Dan Stein, head of the Department of Psychiatry and Director of the Brain-Behaviour Initiative at the University of Cape Town, and director of the South African Medical Research Council's Anxiety and Stress Disorders Research Unit. Together with Prof Odile van den Heuvel of the Vrije Universiteit in Amsterdam, Stein co-leads ENIGMA's efforts on obsessive-compulsive disorder, a psychiatric illness whose basis in the brain is one target of ENIGMA's studies; and he and his Groote Schuur Hospital colleagues, such as Prof John Joska, head of neuropsychiatry, also participate in its work on HIV and substance use, key illnesses in the South African context. "Risk factors for mental illness are complex, so worldwide screening of brain scans and genetic data will help us develop better assessment measures, and understand when treatments are most likely to work," Stein says.
"By relating genetic variations to differences in the living brain, ENIGMA has already identified differences in our DNA that affect the brain,s memory systems, across the world," says Nick Martin, co-founder of the effort in Queensland, Australia, and a former president of the Behavioural Genetics Association. "We can now apply the full power of modern genetic research to tens of thousands of brain scans, giving us new power to understand the causes of brain disease."
The economic and personal cost of brain diseases, such as multiple sclerosis and ALS, is increasing, but many neurological diseases remain hard to treat, as their underlying causes are unknown. The ENIGMA project identifies new sources of disease risk by screening brain scans and genetic tests collected across the world. Currently, computing facilities worldwide analyse ENIGMA's data around the clock, to detect effects of treatments or risk factors, effects that may vary worldwide.
The announcement of funding for ENIGMA comes as part of a major US programme to support 11 national centres of excellence, part of the NIH Big Data to Knowledge Initiative announced in 2013, to discover patterns in large-scale collections of medical data. The efforts targeting large-scale biomedical data promise to discover better diagnostic tools for dementia, schizophrenia, and developmental disorders such as autism, which have been challenging to treat as their root causes are unknown.
"Worldwide efforts in medical research tend to pay off economically," says Michael Weiner, a professor of neurology at the University of California, San Francisco, who leads international efforts to understand Alzheimer's disease. "As more medical tests and scans are collected, concerted efforts from scientists worldwide are needed to make the most of all the data."
 This work is licensed under a Creative Commons Attribution-NoDerivatives 4.0 International License.
This work is licensed under a Creative Commons Attribution-NoDerivatives 4.0 International License.
Please view the republishing articles page for more information.










